OSIRIX LITE SEGMENTATION SERIES
(In Horos, had the same problem with the time series example linked to the dcm image files all in one directory seems to be misinterpretedĪs one big series with multiple mouse heads stacked together). Is just intepreted as a really big stack,īut maybe some other steps are needed to treat/mark as 4D.Ĭlicking the “4D Viewer” icon says that multiple series takenĪt different times are needed as input (whereas my example series that isĦ00. My example time series (subset of mouse brain astrocytoma MRI from TCIA Youtube video not in English but shows some of this stuff. It can be exported to a DICOM file, but again, it's just one 2D image.Ĭan mask data with the 3D segmentation ( ROI. Saving lists the image in the database window from where Thinks a while and then shows a results panel with options including opacityĪnd “Save as DICOM” but unfortunately that only refers to a That goes with the lung dataset) that can be read back into Horos. Region (2D/3D Segmentation) and save it as a. I can create a segmentation using the 2D viewer menu ROI. Import a single segmentation file, nothing seems to happen. Stack, I only see the image stack in the database, and even if I explicitly If I import a directory with both segmentation(s) and a corresponding image Single-DICOM-file segmentations seem to be ignored
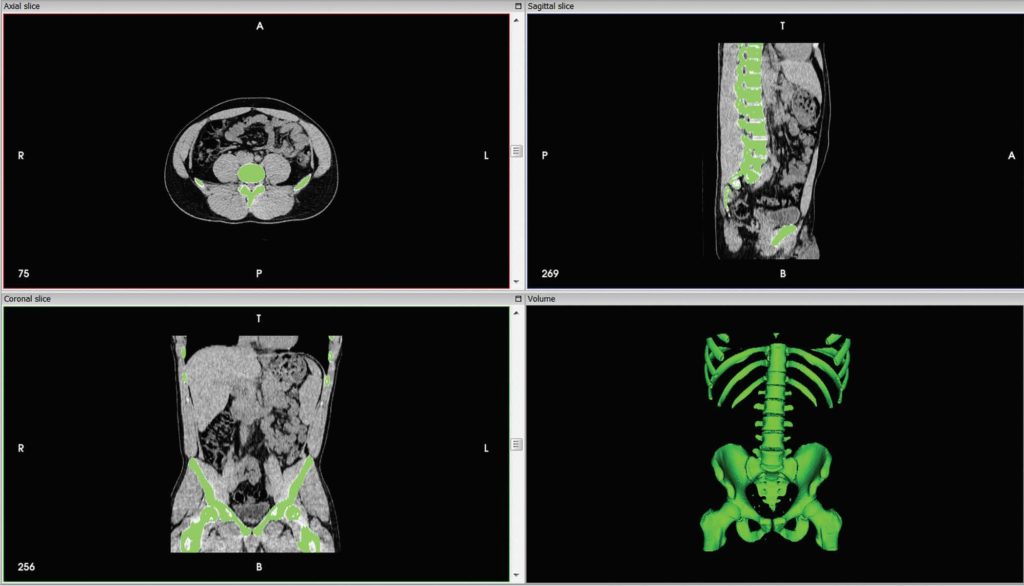
Projection mode, background color, shading, and filtering. Includes not just a CLUT but other settings like Level of detail, shading, parallel (orthographic) vs.
OSIRIX LITE SEGMENTATION REGISTRATION
